The Sequencing-by-Synthesis method, also known as Solexa-Illumina-method, is the most widespread method among all Next-Generation-Sequencing methods (NGS). It was used to identify the RNA fingerprint of the SARS-COV2 virus in January 2020 in a research institute in Wuhan. Since then, this RNA fingerprint measurement has frequently been repeated in research institutes all over the world. And today this NGS method is used worldwide day-by-day to continuously monitor the SARS COV2 virus for mutations. Sub-systems for motion and positioning of imaging optics and flow cells play a decisive role in this method.
XYZ Optical Image Scanning
In addition to microfluidic flushing and washing cycles, as also used by the other NGS methods, e.g. the semiconductor-method and the nanopore-method, the Sequencing-by-Synthesis method is based on very precise and fast XYZ-optical image scanning of the whole sample area.
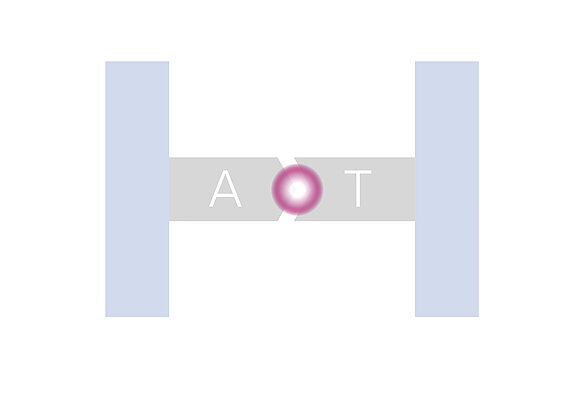
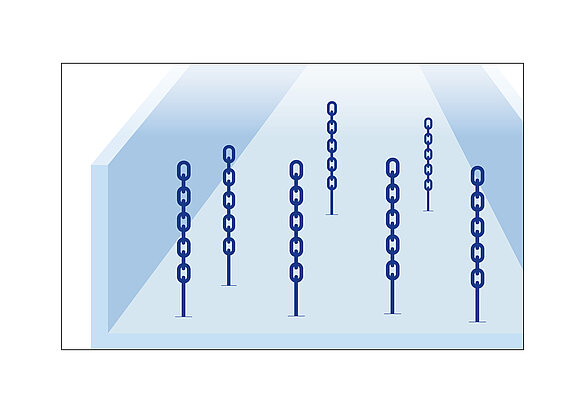
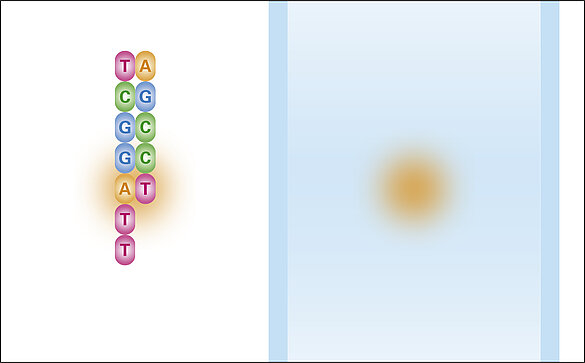
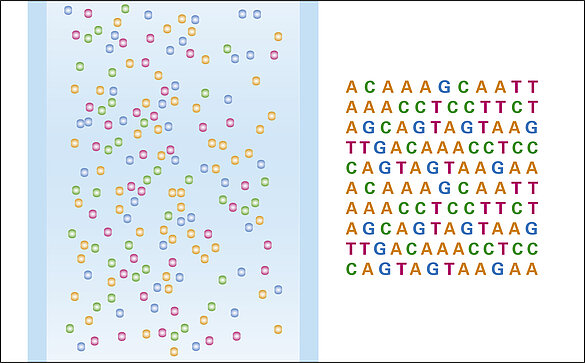
In advance to flushing the flow cells, the whole genome is put to many shorter pieces by the shot-gun technique. These pieces are then PCR amplified and split to single strands (oligo strands) before they are finally attached to the bottom of a flow cell channel. From here the stepwise recombination of base pairs starts vertically and each recombination is indicated by a fluorescence signal. The base pairs contain four different nucleotides to result in four different fluorescence colours.
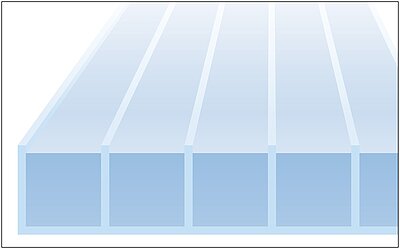
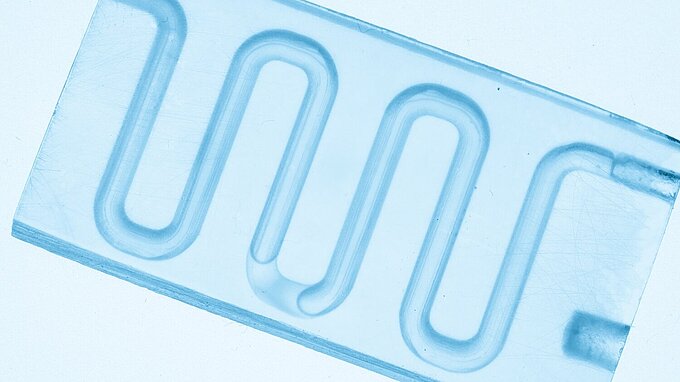
To reduce turbulence and to achieve laminar flows at high speed flushing and washing cycles, flow cells with several parallel channels are used. Typical channel width is some millimetres. This microfluidic pumping and sucking can be done fast, precise and reliable by piezo-based actuators in form of membrane pump plates, benders or tubes.
After each nucleotide flushing the read-out of the whole flow cell has to be done by a XY-scan with all four laser excitation wavelengths. For this purpose a fast and precise motion drive combination for the laser excitation and read-out in every XY-position and at the correct Z-level is needed. Comprising a Z focus adjustment in every XY-coordinate and then scanning the XY-plane typically in a meander-like style.
Motion Solutions from PI
For the XYZ-scanning three drives are needed and typically in addition a rotary drive to correctly position the whole flow cell after cell exchange in the device. This means, in total four precision drives are needed:
Piezo Actuators from PI
Piezo actuators operate at frequencies of up to several kHz and are ideally suited for milli-, micro- and nanodispensing applications: They can switch valves directly, work against a closing spring or a flexible tube for volume displacement and propel liquids within a microfluidic reaction volume – like a membrane pump.